HDU1080(DP)
我用的dp是n^3的, dp[i][j] 表示在s串的i个前和t串的j个前,s[i],t[j]为最末端的两个串得到的最大值。
状态转移方程为:
之前将s和t串最尾端添加'-'
for(int i=;i<=n;i++)
{
for(int j=;j<=m;j++)
{
if(i==n&&j==m) break;
int tmp=;
for(int k=j-;k>=;k--)
{
dp[i][j]=max(dp[i][j],dp[i-][k]+tmp);
tmp += fuc('-',t[k]);
}
tmp=;
for(int k=i-;k>=;k--)
{
dp[i][j]=max(dp[i][j],dp[k][j-]+tmp);
tmp += fuc('-',s[k]);
}
dp[i][j]+=fuc(s[i],t[j]);
}
}
好的一种做法是n^2的dp,dp[i][j]表示s串的i个前,和t串的j个前,得到的最大值,其中s[i],t[j]并不一定要是最末端的元素.
状态转移方程是:
dp[i][j]=max(dp[i-1][j-1]+cos(s[i],t[j]),max(dp[i-1][j]+cos('-',s[i]),dp[i][j-1]+cos('-',t[j])));
Human Gene Functions
Time Limit: 2000/1000 MS (Java/Others) Memory Limit: 65536/32768 K (Java/Others)
Total Submission(s): 1752 Accepted Submission(s): 988
A human gene can be identified through a series of time-consuming biological experiments, often with the help of computer programs. Once a sequence of a gene is obtained, the next job is to determine its function. One of the methods for biologists to use in determining the function of a new gene sequence that they have just identified is to search a database with the new gene as a query. The database to be searched stores many gene sequences and their functions – many researchers have been submitting their genes and functions to the database and the database is freely accessible through the Internet.
A database search will return a list of gene sequences from the database that are similar to the query gene. Biologists assume that sequence similarity often implies functional similarity. So, the function of the new gene might be one of the functions that the genes from the list have. To exactly determine which one is the right one another series of biological experiments will be needed.
Your job is to make a program that compares two genes and determines their similarity as explained below. Your program may be used as a part of the database search if you can provide an efficient one.
Given two genes AGTGATG and GTTAG, how similar are they? One of the methods to measure the similarity of two genes is called alignment. In an alignment, spaces are inserted, if necessary, in appropriate positions of the genes to make them equally long and score the resulting genes according to a scoring matrix.
For example, one space is inserted into AGTGATG to result in AGTGAT-G, and three spaces are inserted into GTTAG to result in –GT--TAG. A space is denoted by a minus sign (-). The two genes are now of equal length. These two strings are aligned:
AGTGAT-G
-GT--TAG
In this alignment, there are four matches, namely, G in the second position, T in the third, T in the sixth, and G in the eighth. Each pair of aligned characters is assigned a score according to the following scoring matrix.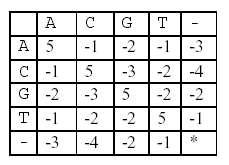
* denotes that a space-space match is not allowed. The score of the alignment above is (-3)+5+5+(-2)+(-3)+5+(-3)+5=9.
Of course, many other alignments are possible. One is shown below (a different number of spaces are inserted into different positions):
AGTGATG
-GTTA-G
This alignment gives a score of (-3)+5+5+(-2)+5+(-1) +5=14. So, this one is better than the previous one. As a matter of fact, this one is optimal since no other alignment can have a higher score. So, it is said that the similarity of the two genes is 14.
7 AGTGATG
5 GTTAG
7 AGCTATT
9 AGCTTTAAA
21
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <algorithm>
#include <math.h>
#include <map>
#include <queue>
#include <sstream>
#include <iostream>
using namespace std;
#define INF 0x3fffffff
#define N 110 int n,m;
int dp[N][N];
char s[N];
char t[N]; int fuc(char x,char y)
{
if(x==y) return ;
if( (x=='-'&&y=='A') || (y=='-'&&x=='A') ) return -;
if( (x=='-'&&y=='C') || (y=='-'&&x=='C') ) return -;
if( (x=='-'&&y=='G') || (y=='-'&&x=='G') ) return -;
if( (x=='-'&&y=='T') || (y=='-'&&x=='T') ) return -;
if( (x=='A'&&y=='C') || (y=='A'&&x=='C') ) return -;
if( (x=='A'&&y=='G') || (y=='A'&&x=='G') ) return -;
if( (x=='A'&&y=='T') || (y=='A'&&x=='T') ) return -;
if( (x=='C'&&y=='G') || (y=='C'&&x=='G') ) return -;
if( (x=='C'&&y=='T') || (y=='C'&&x=='T') ) return -;
if( (x=='G'&&y=='T') || (y=='G'&&x=='T') ) return -;
}
int main()
{
//freopen("//home//chen//Desktop//ACM//in.text","r",stdin);
//freopen("//home//chen//Desktop//ACM//out.text","w",stdout);
int T;
scanf("%d",&T);
while(T--)
{
scanf("%d%s",&n,s+);
s[++n]='-';
scanf("%d%s",&m,t+);
t[++m]='-';
for(int i=;i<=n;i++)
for(int j=;j<=m;j++)
{
dp[i][j] = -INF;
}
dp[][]=;
// dp[i][j] 表示在s串的i个前,t串的j个前,并且s[i]和t[j]在最尾端,
for(int i=;i<=n;i++)
{
for(int j=;j<=m;j++)
{
if(i==n&&j==m) break;
int tmp=;
for(int k=j-;k>=;k--)
{
dp[i][j]=max(dp[i][j],dp[i-][k]+tmp);
tmp += fuc('-',t[k]);
}
tmp=;
for(int k=i-;k>=;k--)
{
dp[i][j]=max(dp[i][j],dp[k][j-]+tmp);
tmp += fuc('-',s[k]);
}
dp[i][j]+=fuc(s[i],t[j]);
}
}
printf("%d\n",max(dp[n][m-],max(dp[n-][m-],dp[n-][m])));
}
return ;
}
最新文章
- T-SQL Transact-SQL 编程
- [LintCode] Reverse Nodes in k-Group 每k个一组翻转链表
- php——用for循环打印半金字塔、金字塔、正方形、倒金字塔、菱形、空心图形等
- 关于div居中
- 修改Fedora 20 启动项
- 初探 FFT/DFT
- 使用ConfuserEx加密混淆程序以及如何脱壳反编译
- [NOIp 2017]列队
- 携程实时计算平台架构与实践丨DataPipeline
- Codeforces 40E Number Table - 组合数学
- css设计技巧
- 单机闭环 使用Nginx+Lua开发高性能Web应用
- WEB-WELCOME-FILE-LIST
- bzoj 3674 可持久化并查集加强版——可持久化并查集
- uml各类图
- SpringBoot日记——Cache缓存篇
- [Node.js]32. Level 7: Working with Lists -- Redis
- Thunder团队第三周 - Scrum会议1
- 导出成可运行jar包时所遇问题的解决办法 - 转载
- SpringCloud02 Eureka知识点、Eureka服务端和客户端的创建、Eureka服务端集群、Eureka客户端向集群的Eureka服务端注册